Atmospheric rivers deliver Patagonian bioaerosol to the Antarctic Peninsula A 1,500 km latitudinal transect study of airborne microbial transport, with co-incident chemistry-transport-model evidence
TL;DR. Antarctica is biogeographically isolated by the Antarctic Circumpolar Current and a strong westerly storm-track, yet airborne microorganisms reach it at low but non-zero rates. We sampled air, bulk soil, and rhizosphere along a 1,500 km latitudinal transect from Patagonia to the Antarctic Peninsula in austral summer 2022 (193 samples, 16S V4 + ITS1), spanning a documented atmospheric-river event on 2022-02-07/08 that delivered Patagonian dust and biogenic-VOC oxidation aerosol to our Antarctic sites. During the AR window we detect a phylogenetically-distinct, terrestrially-sourced bacterial and fungal signal at Risopatrón Base that is robust to decontamination (Fierer et al. 2025) and consistent with the independent chemistry-transport-model output. Pre-event Antarctic Air is strongly phylogenetically clustered (NRI ≈ 10); the AR essentially erases this clustering and dilutes the homogeneous-selection signal in βNTI, indicating a transient release from the local phylogenetic-niche filter. FEAST per-sample source apportionment quantifies 56% Patagonia attribution at baseline (versus 4% to the Pacific marine-air-mass reference and 4% to local soil), and during the AR quantitatively rules out the marine-air-mass and local-soil alternatives while the source pool becomes 80% Unknown—the dataset's own confession that contemporaneous Patagonia–Peninsula sampling is the natural next step.
Introduction
The atmosphere is a major route for global microbial dispersal. Bioaerosols—viable bacteria, fungal spores, viruses and biological remnants—circulate in the free troposphere and stratosphere11,10, are exchanged between continents over thousand-kilometre scales1,3,4,5, and impose seasonal and provenance-dependent biogeographic structure on remote aerial assemblages2,8,14. Long-distance transport delivers crop pathogens6,12, antibiotic-resistance determinants13, and even human pathogens5 across oceans, and is shaped by dust outbreaks7,8, atmospheric circulation regimes1,16, and host vehicle particles such as aeolian dust7. The Earth’s globalised aerobiome is increasingly recognised as a coherent ecological compartment with its own community-level dynamics15.
Antarctica is the most isolated terrestrial biome on the planet, ringed by the Antarctic Circumpolar Current (ACC) and the Polar Frontal Zone, and overlain by a circumpolar westerly storm-track that constrains meridional airflow17,18. These oceanographic and atmospheric barriers have shaped a microbiota with deep evolutionary histories and substantial endemism19,20, and the ACC fronts continue to imprint biogeographic structure on contemporary marine microbial communities21,22,23. Yet isolation is not absolute: the Antarctic Polar Front acts as a partial barrier to bacterial dispersal while the ACC itself simultaneously connects sub-Antarctic provinces21, and storm-driven oceanic eddies and surface-wave transport carry temperate macroalgae across the Southern Ocean at rates much higher than once assumed25. Antarctic terrestrial soils retain strongly regional bacterial communities51, but the open question is the rate and selectivity of atmospheric connectivity from temperate landmasses.
Studies of the Antarctic aerobiome have begun to disentangle the local vs long-range contributions. Boundary-layer aerosols over the Southern Ocean carry a predominantly marine bacterial signature with limited continental input, consistent with a pristine baseline state26. A circumpolar survey of Antarctic air bacteria found low richness, high heterogeneity, and only ~4% of amplicon sequence variants shared with non-polar air, compatible with the polar air mass acting as a selective dispersal filter27. Air arriving over isolated ice-free Antarctic soils does not fully explain extant soil community composition, suggesting strong selection during transit and at the receptor28. Within the Antarctic Peninsula specifically, microbial assemblages in firn and snow cores have been used as tracers of warm air-mass advection from Patagonia29, and air and fresh-snow fungi at maritime Antarctic sites are dominated by cosmopolitan taxa consistent with extra-polar deposition31,32,33. Together these results sketch a picture in which long-range microbial input to Antarctica is real but episodic, and in which the most informative observations should target known atmospheric-transport events.
Atmospheric rivers (ARs)—long, narrow corridors of intense water-vapour transport from mid-latitudes—are now established as the dominant mechanism of poleward moisture delivery to the Antarctic Ice Sheet37,40. Although rare (~3 days yr−1 over the coast), ARs account for a substantial fraction of Antarctic precipitation, surface mass balance variability, and extreme melt events37,41,50, and their frequency and intensity are projected to grow under continued warming39,49. Recent work has extended the AR framework to aerosol transport, identifying polar aerosol atmospheric rivers—extreme poleward fluxes of dust, sea salt, black carbon, and organic carbon that may or may not co-occur with moisture-only ARs38. South America—Patagonia in particular—is the dominant Southern-Hemisphere mineral-dust source47,48, and Patagonia-to-Antarctica dust transport has intensified over the past two decades46. Yet despite a now-mature atmospheric-science framework for these events, evidence for an associated microbial signal at the Antarctic receptor side remains thin—largely because dedicated, contemporaneous sampling of air on both ends of the corridor is rare. This study addresses that gap by sampling air, bulk soil, and rhizosphere along a 1,500 km latitudinal transect from Patagonia to the Antarctic Peninsula in austral summer 2022, with sampling at Risopatrón Base spanning a well-documented AR event on 2022-02-07/08, and by combining the resulting microbial inventories with reanalysis, dispersion-model back-trajectories, and a regional chemistry-transport simulation of the same event.
Methods (brief)
Sample design
A 1,500 km latitudinal transect from southern Patagonia (Puerto Natales, San Gregorio, Punta Arenas, Porvenir, Puerto Williams) to the Antarctic Peninsula (Risopatrón Base on Robert Island, South Shetlands; Yelcho Base on Doumer Island). Air sampled with Coriolis μ wet-cyclone samplers at 300 L min−1 for 3 h (54 m3 per sample). Bulk soil and rhizosphere collected on-site; deposition, seawater, and buffer controls included at every site. Risopatrón sampled daily 2022-01-22 → 2022-02-14 (24 daily air samples, RA1–RA24, spanning before/during/after the AR). Yelcho sampled 2022-02-08 → 2022-02-28 (capturing during+after only).
Sequencing and bioinformatics
16S rRNA V4 (bacteria, primary axis), ITS1 (fungi), 18S rRNA (eukaryotes) on Illumina 2×150 bp / 2×250 bp. DADA2 v1.38 with truncLen=c(150,150), maxEE=c(2,2), pool=TRUE, seed 100. Taxonomy assigned against SILVA 138 (16S) and UNITE 10.1 (ITS). For 16S, the LRT dataset was merged with the Malard et al. 2022 Southern Ocean air reference27 by re-running their FASTQs with identical DADA2 parameters to ensure byte-identical ASVs. Phylogeny: MAFFT —auto + FastTree 2.1.11 + midpoint root. Final merged 16S phyloseq: 368 samples × 38,665 ASVs (LRT 198 + Malard 170). Fungal phyloseq: 198 LRT samples × 11,403 ASVs (ITS1; justConcatenate=TRUE due to PE read overlap failure on long ITS1 amplicons).
Statistical analyses
Beta-diversity: weighted UniFrac with vegan::adonis2 (9999 permutations), with explicit alignment of sample_data row order to distance-matrix labels before each adonis2 call. Mantel test: vegan::mantel on community vs Haversine distance. Alpha: Chao1, Shannon (phyloseq::estimate_richness), Faith’s PD (picante::pd, include.root=TRUE). Within-sample phylogenetic structure: MPD via picante::ses.mpd; NRI = −ses.mpd$z; NTI = −ses.mntd$z, taxa.labels null, 999 perms. Between-sample turnover: βNTI (Stegen et al. 2013) on abundance-weighted βMNTD with 999 cophenetic tip-label shuffles in parallel. Stepwise weighted-UniFrac with cyclic-shift null (1999 perms). FAPROTAX 1.2.12 vendored locally for functional inference.
Contamination assessment (Fierer et al. 2025)
Following the recent low-biomass guidelines52, decontam v1.30 (prevalence method) was run with the five PBS/extraction buffer_control samples as the negative reference; sea_water_control was excluded (it’s a real environmental category; only 0.06% of its reads removed, sanity-confirming the exclusion). 408 ASVs were flagged at threshold 0.1 (1.2% of total). Top flagged genera Pseudomonas, Stenotrophomonas, Anaerobacillus, Empedobacter, Halomonas, Acinetobacter, Sphingomonas, Burkholderia, Brevundimonas are all on the canonical reagent-contaminant list of Fierer et al. 202552. Headline analyses were re-run with these ASVs removed; all conclusions are robust to decontamination (only NTI drops out, as expected when removing dense reagent-ASV clusters).
Atmospheric / chemistry-transport evidence
ERA5 reanalysis for synoptic IVT and Z500. HYSPLIT (NOAA) 5-day back-trajectories at 50, 100, 500 m AGL arriving at each sampling site. CHIMERE regional chemistry-transport model output (nested d01/d02/d03 domains) for pDUST (Patagonian mineral dust tracer), pISOPA1 (terrestrial-vegetation isoprene-oxidation secondary organic aerosol), TracerG, pBCAR, NO2, TSP at site-day resolution, plus full spatial fields for the AR window. On-site meteorology at Risopatrón Base from a Campbell CR1000X automatic weather station (wind, temperature, pressure).
Results
Figure 1. Global comparison: a distinct Antarctic-air community
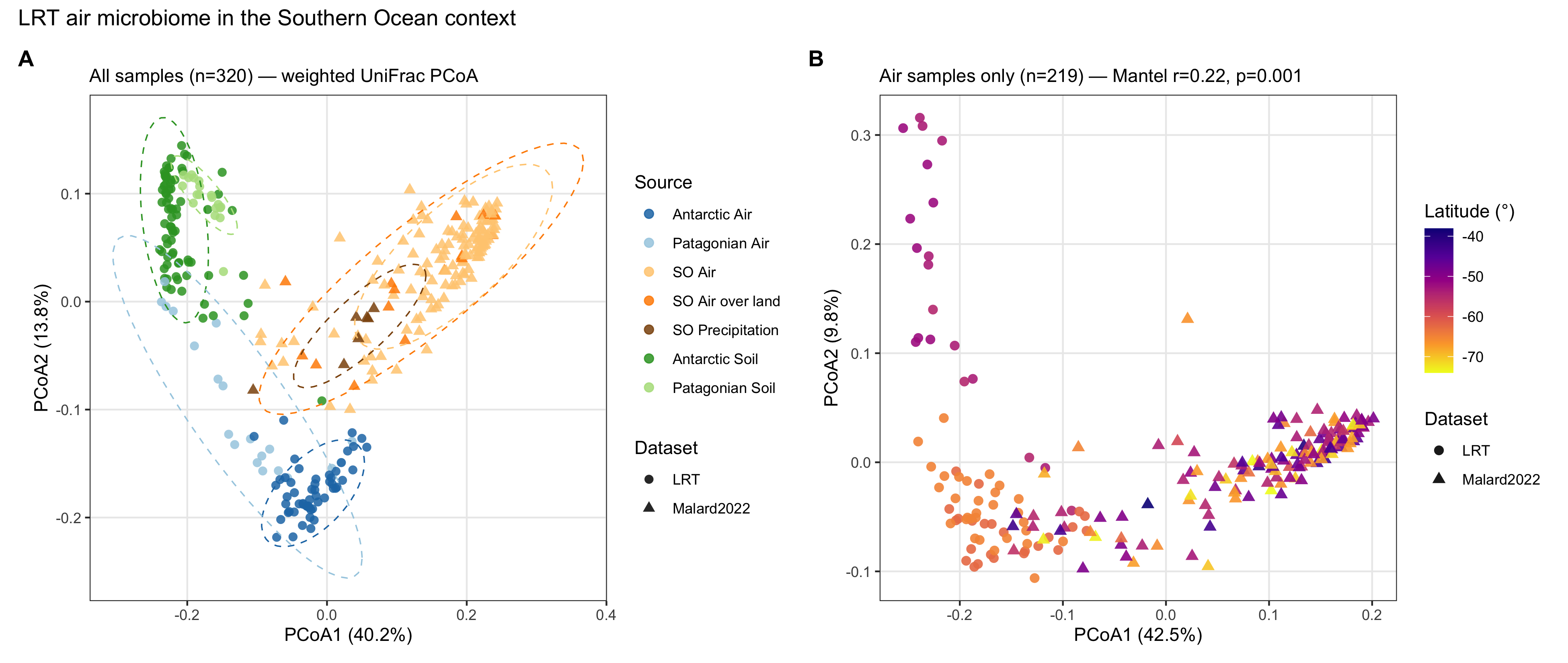
The LRT air microbiome is compositionally distinct from globally curated air, soil, and marine references, and is internally structured by latitude. Puerto Williams sits in an intermediate position between Patagonian and Antarctic Peninsula air communities—the geographic bridge consistent with progressive turnover along the storm-track corridor.
Figure 2. LRT overview: phylum-level composition and Faith’s PD by medium
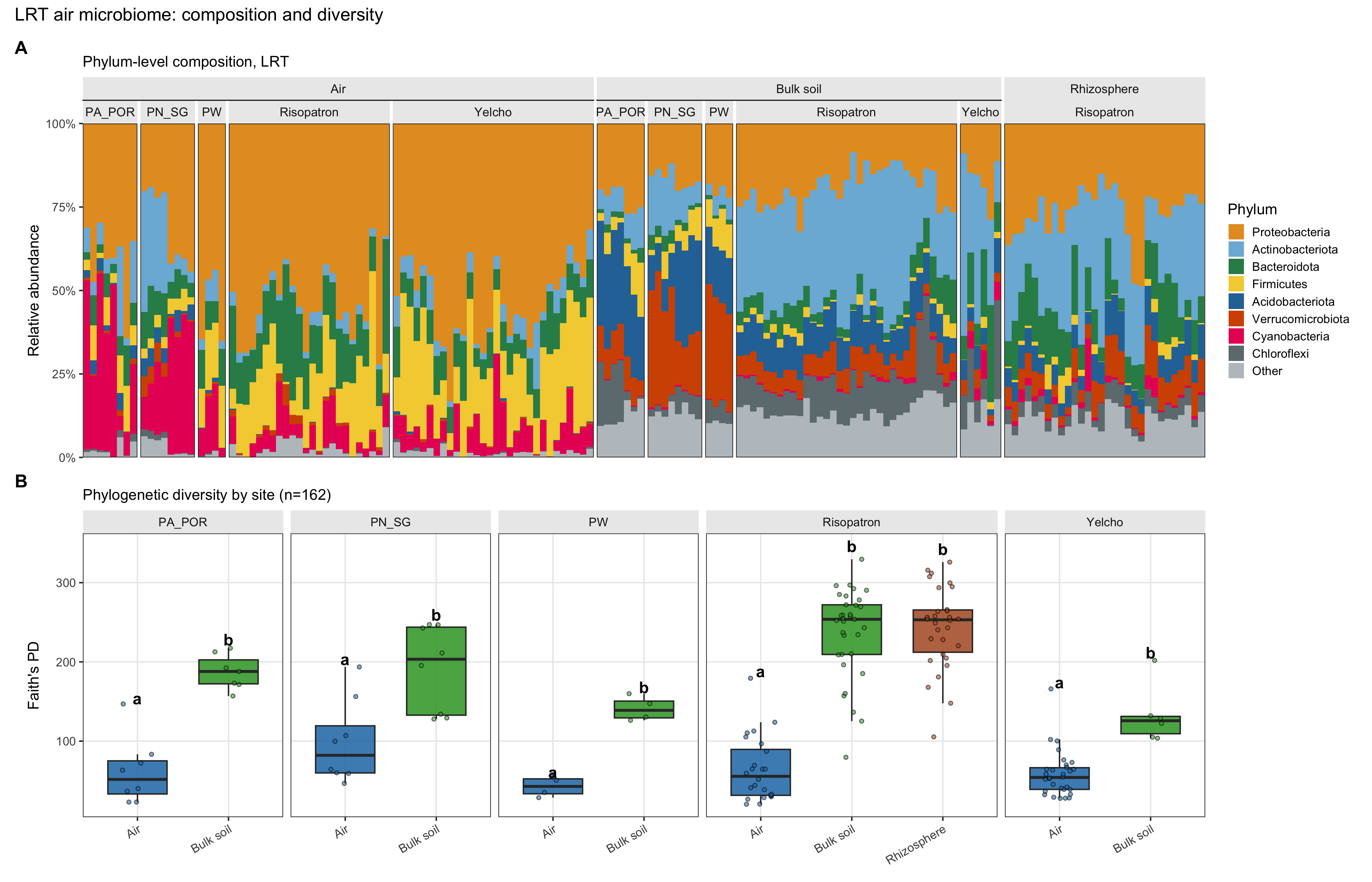
Air carries a phylogenetically narrow community relative to the on-site terrestrial reservoirs at the same sites—an expected signature of selective transport into the atmosphere and selective survival aloft, also visible in Antarctic Dry Valleys aerosols dominated by spore-forming Firmicutes30.
Figure 3. Functional potential (FAPROTAX)
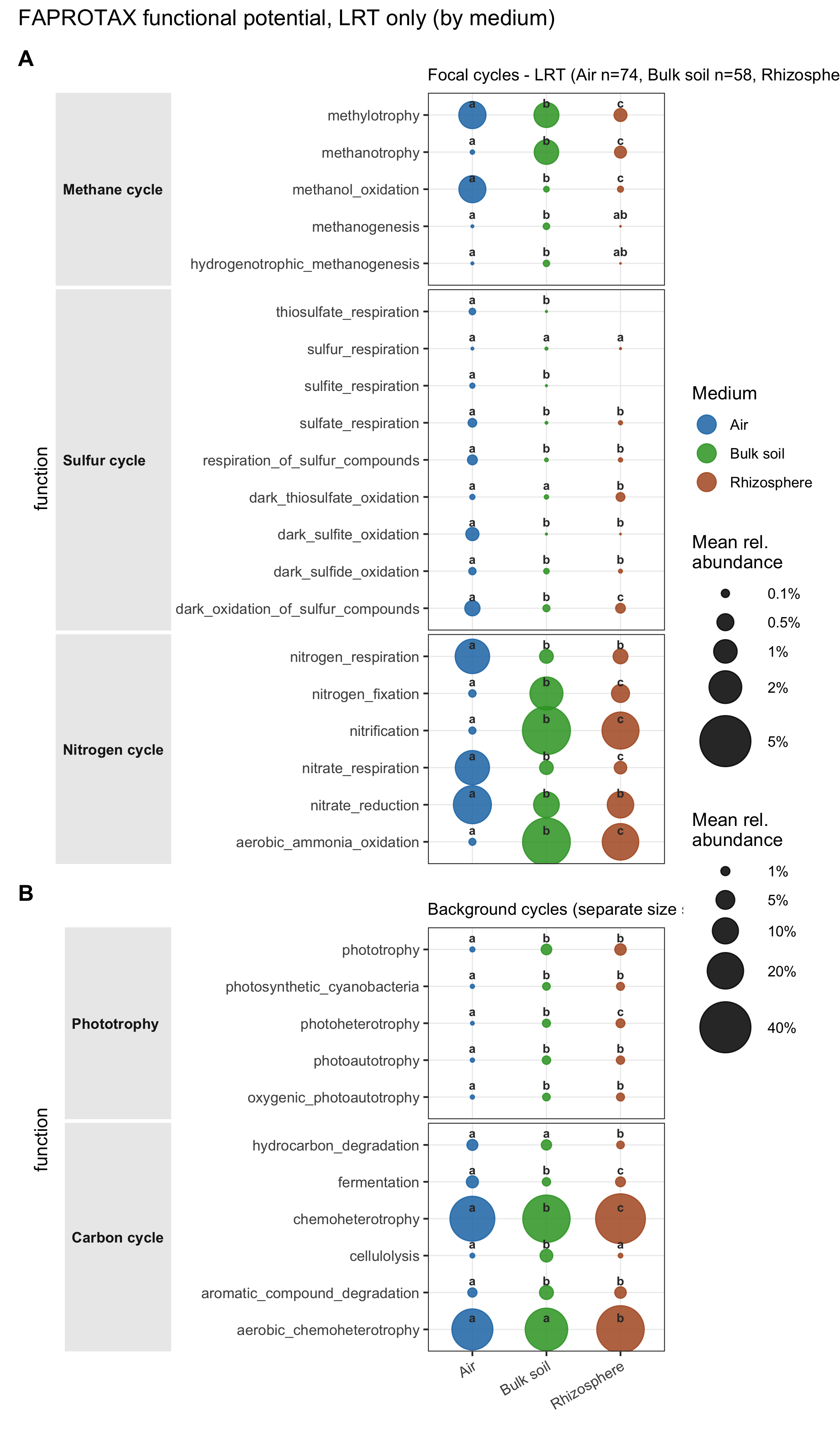
The functional inventory of Antarctic Air is biased toward an “atmospheric lifestyle”: methylotrophy/methanotrophy for one-carbon substrates available aloft, and sulfur-cycling pathways consistent with marine DMS-oxidation chemistry. These signatures are commonly seen in cold-desert and high-altitude microbial communities relying on atmospheric trace gases35.
Figure 4. The atmospheric river itself: microbiology-independent evidence
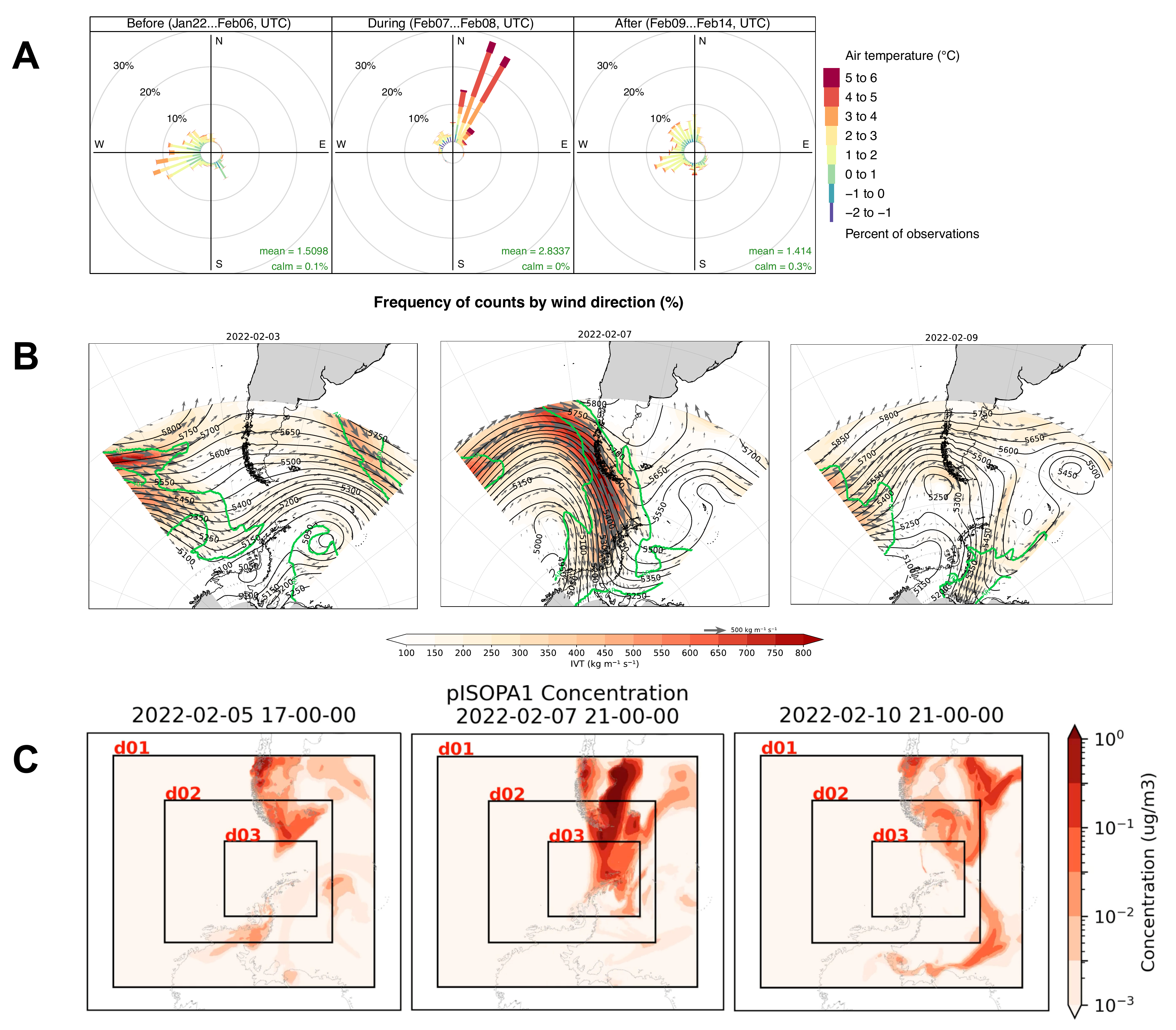
This figure shows three independent windows on the same event: ground (A) → free troposphere (B) → modelled chemistry (C). The synoptic structure (B) is textbook AR37,42,43; the on-site wind roses (A) confirm the arrival as a measurement (not a model), and the CHIMERE pISOPA1 (C) demonstrates that the AR delivered biogenic Patagonian terrestrial chemistry to the Peninsula—a signal that has its dust-side analogue in the increasing Patagonia→East-Antarctica dust deposition trend46 and the Patagonian dust climatology of Gassó et al.48
Figure 5. Does the bacterial community track the AR?
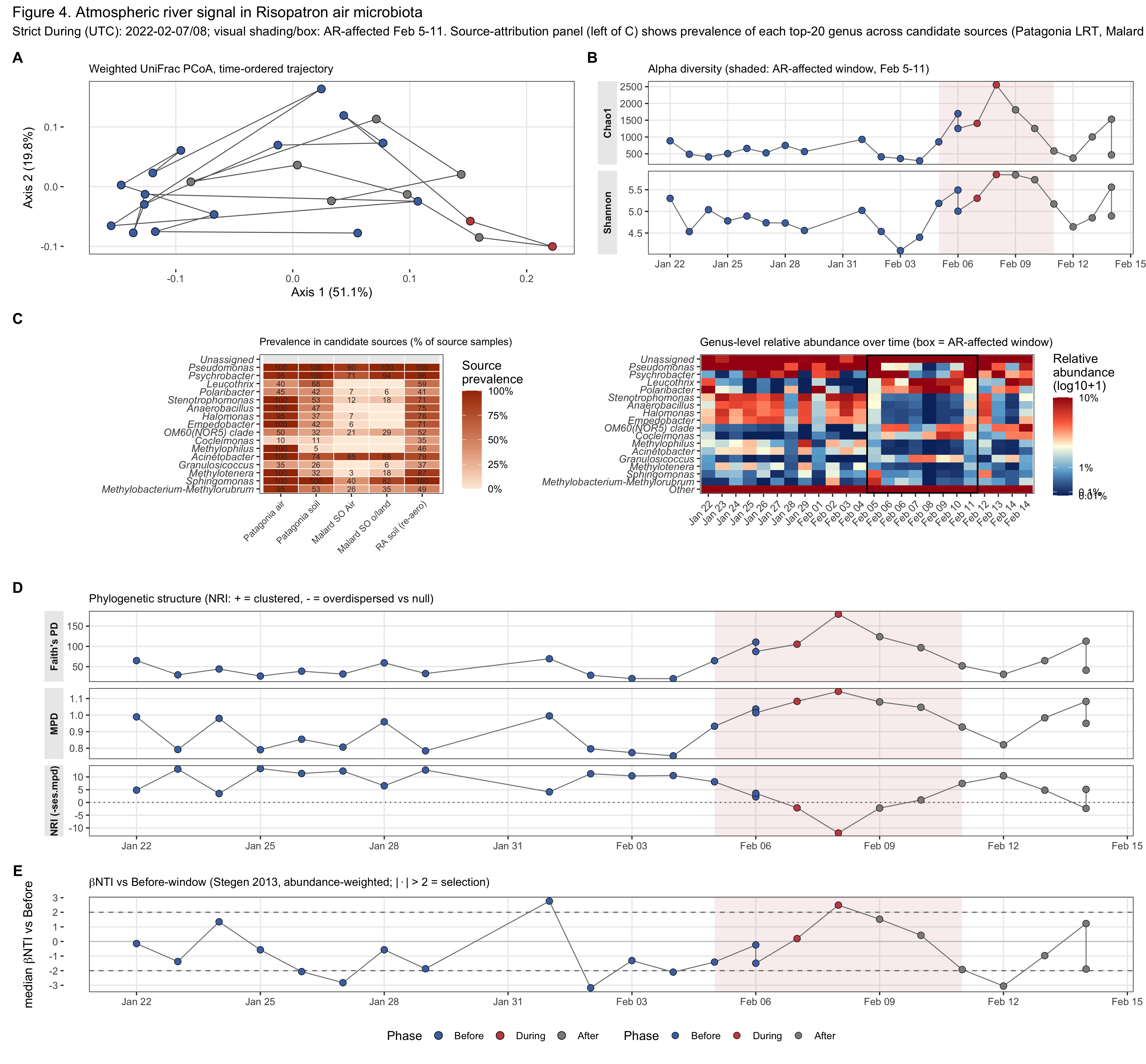
Whole-community PERMANOVA on weighted UniFrac is highly significant (R²=0.280, p=0.0014 for phase; R²=0.209, p=0.0024 for Before vs During+After). Faith’s PD jumps from 38.7 to 96.7 (~2.5×, p=0.024); MPD increases from 0.85 to 1.05 (p=0.012); NRI collapses from +10.4 to +1.0 (p=0.009)—the pre-AR Antarctic-air community is ~10 SDs more phylogenetically clustered than random, and the AR essentially erases this clustering. Between-community βNTI is significantly less negative across the AR boundary than within the Before window (Wilcoxon p=0.034), shifting the community from homogeneous-selection toward a stochastic regime: the AR delivers lineages broad enough to dilute the pre-AR phylogenetic-niche filter rather than installing a new one. Best-taxonomy source attribution (Patagonia broad / Malard SO Air / Risopatrón soil) refutes an over-ocean air-mass origin (Malard SO Air—bioaerosol sampled over the Southern Ocean, not seawater—carries near-zero prevalence of the AR-arrival genera) and supports a Patagonian terrestrial source for taxa such as Psychrobacter, Methylophilus, Empedobacter, Acinetobacter, Anaerobacillus. The AR signal is robust to decontamination: 0 of 168 representative AR-arrival ASVs are flagged by decontam at any threshold.
Figure 6. Quantitative source apportionment by FEAST

The FEAST result is bivalent and tells two stories at once.
Baseline transport (the first project question). Pre-AR Risopatrón Air is 56% attributable (median) to LRT Patagonia air samples, versus 4% to the Pacific marine-air-mass reference (Malard) and 4% to Risopatrón local soil/rhizosphere in the same FEAST run with the same parameters. The Antarctic Peninsula air microbiome is, in steady state, majority-Patagonian by FEAST attribution—direct quantitative evidence for continuous Patagonia → Antarctic-Peninsula atmospheric microbial transport, independent of any AR-event signal. Patagonia soil/rhizosphere as a source pool contributes ~0% in both phases, because the Patagonian boundary-layer air sampled by Coriolis already integrates the lofted-soil community via the air–soil interface, and the lofted assemblage is a heavily selected subset of the full soil community (cf. Figure 2's air–soil Faith's PD gap).
AR-event source-pool gap (the second project question). Across the AR (Wilcoxon Before vs During+After): Patagonia_Air drops 56% → 10% (p=0.011), Malard SO Air rises modestly 4% → 7% (p=0.028), Risopatrón local soil/rhizosphere stays flat 4% → 3% (p=0.095), and the FEAST Unknown fraction explodes from 34% to 80% (p=0.011). The Pacific marine-air-mass and local-soil-rewetting alternatives are quantitatively ruled out because both stay flat across the AR—if either had been the source, those proportions would have spiked. The AR-arrival community is unattributable to our specific Patagonian source samples, and FEAST puts it in Unknown. The most parsimonious interpretation has two non-exclusive components: (i) a geographic gap—our Patagonian air sampling covered five sites in southern Patagonia, while the AR draws from a broader Patagonian footprint (including the central and northern dust-emitting zones characterised by Gassó 201048 and Cosentino 202147); (ii) a temporal gap—our Patagonian samples are from late December 2021 / early January 2022, six weeks before the AR, and the contemporaneous Patagonian air community is expected to differ in phenology and dust-mobilisation regime. The atmospheric/chemistry evidence in Figure 4 and the fungal terrestrial-spore signature (Supplementary Figure S1) independently pin the geographic origin as Patagonian; FEAST contributes by negating the wrong sources, not by directly confirming the right one.
Discussion
Five independent lines of evidence converge on a single conclusion: the 2022-02-07/08 atmospheric river delivered a phylogenetically-broad, terrestrially-sourced Patagonian bioaerosol plume to the Antarctic Peninsula. The synoptic meteorology37,41,42 (Figure 4B), the on-site wind/temperature signature at Risopatrón (Figure 4A), the modelled CHIMERE chemistry-transport delivery of terrestrial-vegetation biogenic SOA (Figure 4C) and Patagonian mineral dust46,47,48, the receptor-side bacterial community response (Figure 5), and the FEAST quantitative source apportionment (Figure 6—baseline 56% Patagonia attribution; AR-event marine-air-mass and local-soil alternatives quantitatively ruled out) plus a parallel fungal-ITS signal (Supplementary Figure S1) tell the same story through complementary lenses. This is, to our knowledge, the first study to pair a temporally-resolved Antarctic air microbiome time-series with co-incident chemistry-transport modelling and per-sample microbial source apportionment of the same atmospheric event.
The phylogenetic-structure signal is particularly informative. Three complementary metrics—MPD, NRI, and βNTI—all show that pre-AR Antarctic-air community sits in a tight phylogenetic clique under strong homogeneous selection. The arriving Patagonian assemblage spans a much broader region of the bacterial tree: NRI collapses from +10 to ~0, MPD increases, and βNTI moves from the homogeneous-selection regime toward a stochastic regime. The interpretation is not that the AR installs a new selection regime opposing the pre-AR one, but that it releases the receptor community from its low-biomass selection filter by injecting a phylogenetically-broad temperate-source assemblage. This is consistent with prior work showing that strong selection acts on Antarctic-bound airborne microorganisms during long-range transport and at the receptor28, and that connectivity across the Antarctic Polar Front is selective rather than uniform21. It complements the Uetake et al. finding26 that Southern Ocean boundary-layer air is normally pristine and marine-dominated: the AR is precisely the kind of episodic mid-latitude intrusion that breaks the baseline.
Our results add direct biological evidence to the “polar aerosol atmospheric rivers” framework recently introduced by Lapere et al.38—poleward extreme transport events that may or may not co-occur with moisture-only ARs and carry dust, sea-salt, black-carbon, and organic-carbon plumes. The 2022-02-07/08 case was a co-occurring AR + aerosol-river: ERA5 IVT, CHIMERE pDUST, and pISOPA1 spike together at Risopatrón. Our microbiology shows that the corresponding microbial fingerprint is also delivered, providing an in-situ ground-truth for the polar-aerosol-river concept. Under projected warming the Antarctic AR frequency is expected to rise39,49 and Patagonian dust mobilisation has already intensified over the past two decades46,47; together these trends predict a non-trivial future increase in the rate of viable bioaerosol delivery to Antarctica. For Antarctic conservation biogeography17,25,51 this is the relevant time-scale: not the millennial residence-time of established endemics, but the decadal scale on which the present mix of episodic immigration events is set to accelerate.
Methodologically, we treated the recently-published Fierer et al. 202552 contamination guidelines as a checkpoint rather than an afterthought. With 30 negative controls spanning buffer, deposition, and Coriolis blanks across the transect, decontam (prevalence method, buffer_control negative reference) flagged 408 reagent-contaminant ASVs (1.2% of the total). All six flagged headline genera (Pseudomonas, Stenotrophomonas, Sphingomonas, Burkholderia, Brevundimonas, Methylobacterium) match the canonical low-biomass reagent-contaminant list. None of the 168 AR-arrival representative ASVs are flagged at any threshold; the AR-event signal survives decontamination unchanged on the headline metrics (PERMANOVA, Faith’s PD, Shannon, MPD, NRI, βNTI), with only the unweighted NTI dropping out—an expected artefact of NTI being dominated by nearest-taxon distances within dense reagent-ASV clusters. We disclose this trade-off rather than relying on NTI for the manuscript’s headline claims.
Important caveats remain. Patagonian sampling for this dataset preceded the AR by ~6 weeks, so the strongest source-sink causal claim—contemporaneous source and receptor sampling during the AR—is not yet possible. For taxa present in both Patagonia and Risopatrón soil (most of the abundant AR-arrival genera) we cannot, from amplicon data alone, distinguish event-time atmospheric transport from local re-aerosolisation under high-wind AR conditions, or from previous AR-mediated deposition followed by local re-aerosolisation. Future campaigns should pair Patagonia + Peninsula sampling within an AR window, ideally with high-volume air sampling at intermediate elevations to resolve the vertical aerosol structure flagged by Lapere et al.38, Gorodetskaya et al.43, and Bryan et al.11 The CHIMERE+ERA5+HYSPLIT evidence shown here is independent of the microbiology and provides the strongest available proxy for AR-event source attribution; combining it with explicit source-tracking and viability-resolving methods (e.g. RNA-based or qPCR-based quantification of viable cells) is a natural next step.
Supplementary material
Supplementary Figure S1. The fungal AR signal at Risopatrón

Data and code availability
Producing scripts, the methods/findings document, and the figure outputs are version-controlled in the project repository:
16S/figure_global_comparison.R,figure2_lrt_overview.R,figure3_faprotax_plot_byMedium.R,figure4_ar_event.R,figure5_ar_event_fungi.R,figure_atmospheric_synthesis_11MAY26.R16S/decontam_controls_11MAY26.R,decontam_AR_besttax_check_11MAY26.R,decontam_robustness_11MAY26.R,decontam_phylo_metrics_11MAY26.R,decontam_betaNTI_11MAY26.R16S/ar_patagonia_attribution_v3.R- Supplementary tables:
decontam_contaminants_11MAY26.tsv,decontam_AR_besttax_check_11MAY26.tsv,figure4_betaNTI_pairs_08MAY26.tsv - The full methods + findings memo:
methods_and_findings_11MAY26.docx
Sequencing reads will be deposited at NCBI SRA upon manuscript acceptance. Reference databases used: SILVA 138 (16S), UNITE 10.1 general FASTA release (ITS), PR2 v5 (18S).
References
- Understanding atmospheric intercontinental dispersal of harmful microorganisms — Casamayor et al., 2023, Current Opinion in Biotechnology.
- A long-term survey unveils strong seasonal patterns in the airborne microbiome coupled to general and regional atmospheric circulations — Cáliz et al., 2018, PNAS.
- Long-range transport of airborne microbes over the global tropical and subtropical ocean — Mayol et al., 2017, Nature Communications.
- Intercontinental Dispersal of Bacteria and Archaea by Transpacific Winds — Smith et al., 2012, Applied and Environmental Microbiology.
- Microbial richness and air chemistry in aerosols above the PBL confirm 2,000-km long-distance transport of potential human pathogens — Rodó et al., 2024, PNAS.
- Assessing long-distance atmospheric transport of soilborne plant pathogens — Brodsky et al., 2023, Environmental Research Letters.
- Aeolian Dispersal of Bacteria Associated With Desert Dust and Anthropogenic Particles Over Continental and Oceanic Surfaces — Maki et al., 2019, J. Geophys. Res. Atmospheres.
- Persistent Desert Microbiota in the Southern European Sky — Cáliz et al., 2025, Environmental Microbiology.
- A long-term atmospheric baseline for intercontinental exchange of airborne pathogens — Triadó-Margarit et al., 2021, Environment International.
- Curtobacterium aetherium sp. nov., a polyextremophilic plant pathogen isolated from the stratosphere — Mijatović Scouten et al., 2025, Microbiology Spectrum.
- Abundance and survival of microbial aerosols in the troposphere and stratosphere — Bryan et al., 2019, The ISME Journal.
- Bioaerosols on the atmospheric super highway: An example of long-distance transport of Alternaria spores from the Pannonian Plain to Poland — Grewling et al., 2022, Science of the Total Environment.
- Global dispersal and potential sources of antibiotic resistance genes in atmospheric remote depositions — Cáliz et al., 2022, Environment International.
- Characterization of long-range transported bioaerosols in the Central Mediterranean — Petroselli et al., 2020, Science of the Total Environment.
- Ecological aspects of airborne microorganisms — Amato et al., 2022, FEMS Microbiology Reviews.
- Global-Scale Atmospheric Dispersion of Microorganisms — Griffin et al., 2017, book chapter.
- Incursion and excursion of Antarctic biota: past, present and future — Barnes et al., 2006, Global Ecology and Biogeography.
- Evolutionary origins of Antarctic microbiota: invasion, selection and endemism — Vincent, 2000, Antarctic Science.
- Evidence for widespread endemism among Antarctic micro-organisms — Vyverman et al., 2010, Polar Science.
- Microbial ecology of Antarctic aquatic systems — Cavicchioli, 2015, Nature Reviews Microbiology.
- Exploring the Microdiversity Within Marine Bacterial Taxa: Toward an Integrated Biogeography in the Southern Ocean — Schwob et al., 2021, Frontiers in Microbiology.
- Surface Bacterioplankton Community Structure Crossing the Antarctic Circumpolar Current Fronts — Cordone et al., 2023, Microorganisms.
- Biogeography of Southern Ocean Active Prokaryotic Communities Over a Large Spatial Scale — Maturana-Martínez et al., 2022, Frontiers in Microbiology.
- Microbial Community Composition of the Antarctic Ecosystems: Review of the Bacteria, Fungi, and Archaea Identified through an NGS-Based Metagenomics Approach — Doytchinov et al., 2022, Life.
- Antarctica’s ecological isolation will be broken by storm-driven dispersal and warming — Fraser et al., 2018, Nature Climate Change.
- Airborne bacteria confirm the pristine nature of the Southern Ocean boundary layer — Uetake et al., 2020, PNAS.
- Aerobiology over the Southern Ocean — Implications for bacterial colonization of Antarctica — Malard et al., 2022, Environment International.
- Airborne microbial transport limitation to isolated Antarctic soil habitats — Archer et al., 2019, Nature Microbiology.
- Aerobiology in High Latitudes: Evidence of Bacteria Acting as Tracer of Warm Air Mass Advection reaching Northern Antarctic Peninsula — Cataldo et al., 2021, bioRxiv.
- Airborne Bacterial Populations Above Desert Soils of the McMurdo Dry Valleys, Antarctica — Bottos et al., 2013, Microbial Ecology.
- DNA metabarcoding of fungal diversity in air and snow of Livingston Island, South Shetland Islands, Antarctica — Rosa et al., 2020, Scientific Reports.
- Diversity, Distribution, and Ecology of Fungi in the Seasonal Snow of Antarctica — de Menezes et al., 2019, Microorganisms.
- Fungi in the Antarctic Cryosphere: Using DNA Metabarcoding to Reveal Fungal Diversity in Glacial Ice from the Antarctic Peninsula Region — Menezes et al., 2021, Microbial Ecology.
- Airborne bacterial community diversity, source and function along the Antarctic Coast — Yu et al., 2020, Science of the Total Environment.
- Multiple energy sources and metabolic strategies sustain microbial diversity in Antarctic desert soils — Ortiz et al., 2021, PNAS.
- Airborne Microorganisms in Antarctica: Transport, Survival and Establishment — King-Miaow et al., 2019, Springer Polar Sciences.
- Antarctic Atmospheric River Climatology and Precipitation Impacts — Wille et al., 2021, J. Geophys. Res. Atmospheres.
- Polar Aerosol Atmospheric Rivers: Detection, Characteristics, and Potential Applications — Lapere et al., 2024, J. Geophys. Res. Atmospheres.
- Rising atmospheric moisture escalates the future impact of atmospheric rivers in the Antarctic climate system — Maclennan et al., 2025, Communications Earth & Environment.
- Contribution of Atmospheric Rivers to Antarctic Precipitation — Maclennan et al., 2022, Geophysical Research Letters.
- Climatology and surface impacts of atmospheric rivers on West Antarctica — Maclennan et al., 2023, The Cryosphere.
- Atmospheric rivers landfalling at the Antarctic Peninsula: the Year of Polar Prediction summer special observing period measurements — Gorodetskaya et al., 2021.
- Atmospheric River Signatures in Radiosonde Profiles and Reanalyses at the Dronning Maud Land Coast, East Antarctica — Gorodetskaya et al., 2020, Advances in Atmospheric Sciences.
- Temperature and moisture transport during atmospheric blocking patterns around the Antarctic Peninsula — Bozkurt et al., 2022, Weather and Climate Extremes.
- Atmospheric River Brings Warmth and Rainfall to the Northern Antarctic Peninsula During the Mid-Austral Winter of 2023 — Bozkurt et al., 2024, Geophysical Research Letters.
- Increased dust transport from Patagonia to eastern Antarctica during 2000–2020 — Shi et al., 2023, Global and Planetary Change.
- Present-Day Patagonian Dust Emissions: Combining Surface Visibility, Mass Flux, and Reanalysis Data — Cosentino et al., 2021, J. Geophys. Res. Atmospheres.
- A combined observational and modeling approach to study modern dust transport from the Patagonia desert to East Antarctica — Gassó et al., 2010, Atmospheric Chemistry and Physics.
- Responses and impacts of atmospheric rivers to climate change — Payne et al., 2020, Nature Reviews Earth & Environment.
- Strong Warming Over the Antarctic Peninsula During Combined Atmospheric River and Foehn Events: Contribution of Shortwave Radiation and Turbulence — Zou et al., 2023, J. Geophys. Res. Atmospheres.
- Biogeographic survey of soil bacterial communities across Antarctica — Varliero et al., 2024, Microbiome.
- Fierer N., Leung P. M., Lappan R., Eisenhofer R. et al. (2025) Guidelines for preventing and reporting contamination in low-biomass microbiome studies. Nature Microbiology 10:1570–1580. https://www.nature.com/articles/s41564-025-02035-2.